|
1/18/2024 0 Comments Okazaki fragments dna ligase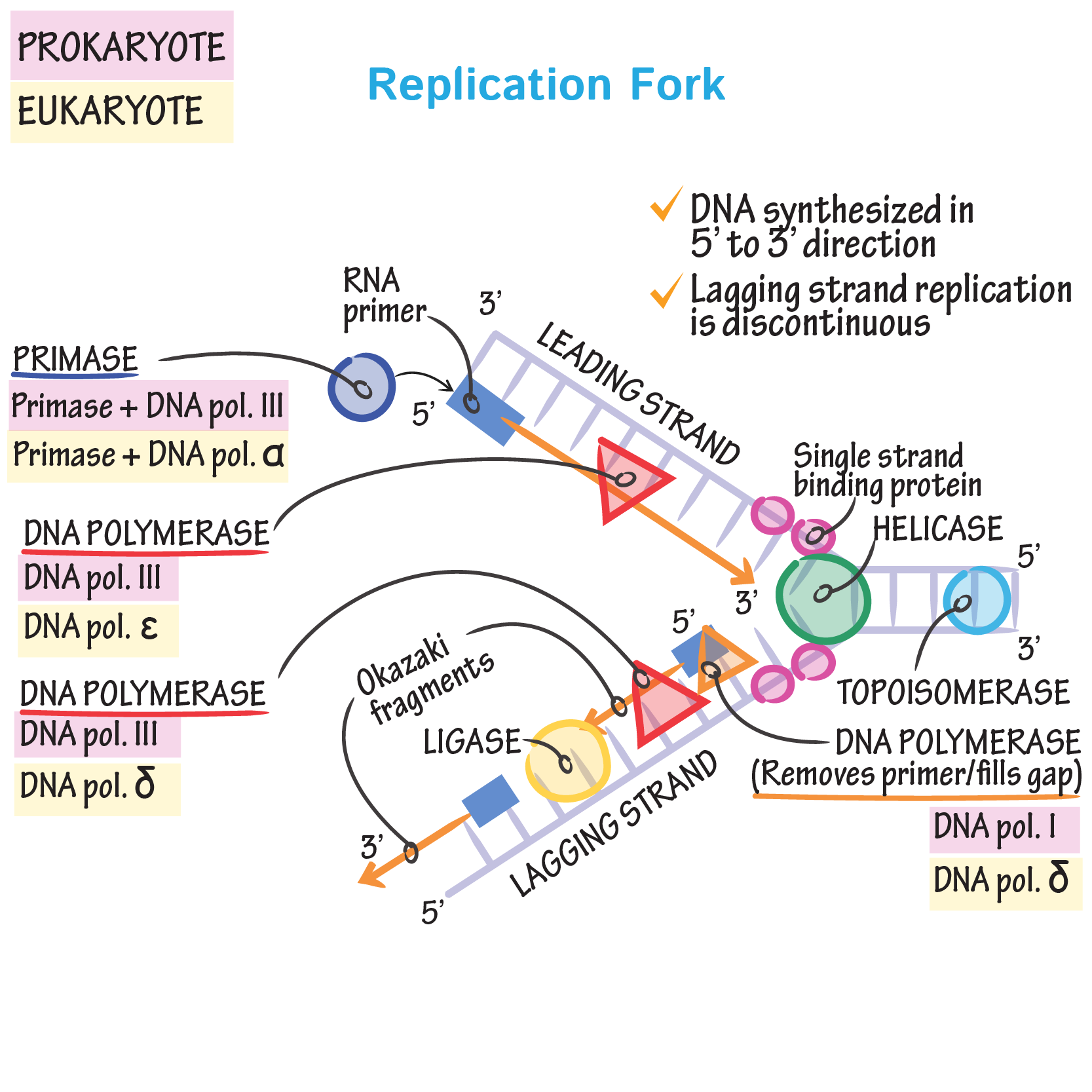
The rate at which these additions are generated increases upon concomitant inactivation of DNA mismatch repair, or by inactivation of the Fen1 Rad27 Okazaki fragment maturation (OFM) nuclease. Here, we show that an engineered low-fidelity LIG1 Cdc9 variant confers a novel mutator phenotype in yeast typified by the accumulation of single base insertion mutations in homonucleotide runs. Whether intrinsic ligation fidelity contributes to the accuracy of replication of the nuclear genome is unknown. 2002 115:1551–1561.DNA ligase 1 (LIG1, Cdc9 in yeast) finalizes eukaryotic nuclear DNA replication by sealing Okazaki fragments using DNA end-joining reactions that strongly discriminate against incorrectly paired DNA substrates. DNA ligase I null mouse cells show normal DNA repair activity but altered DNA replication and reduced genome stability. Enzymes and reactions at the eukaryotic DNA replication fork. RPA governs endonuclease switching during processing of Okazaki fragments in eukaryotes. Dna2 of Saccharomyces cerevisiae possesses a single-stranded DNA-specific endonuclease activity that is able to act on double-stranded DNA in the presence of ATP. Characterization of the enzymatic properties of the yeast Dna2 helicase/endonuclease suggests a new model for Okazaki fragment processing. Black lines represent the DNA template, while pink ones are the newly synthesized DNA and light pink ones are the RNA primers.īae S.H., Seo Y.S. Major proteins factors present in a typical replication fork include: ( i) mini-chromosome maintenance (MCM) proteins (six homo-subunits), which are helicases for opening up the DNA duplex to initiate a DNA replication fork ( ii) RPA, a single-stranded DNA binding protein to protect the DNA template from nuclease cleavage ( iii) primase (a complex of RNA polymerase and Pol α), which synthesizes RNA primers and a short DNA fragment to initiate Okazaki fragments ( iv) Pol δ, the DNA polymerase responsible for synthesizing the major portion of Okazaki fragments ( v) Pol ɛ, the DNA polymerase responsible for leading strand DNA synthesis ( vi) PCNA, which is the DNA clamp for the processivity of DNA polymerase and coordination of Okazaki fragment maturation processes ( vii) RFC, which is the clamp loader for PCNA to load onto DNA duplex ( viii) nucleases, including RNase H, DNA2 and FEN1 for removal of RNA primers and ( ix) DNA Lig I, which joins processed Okazaki fragments into an intact DNA lagging strand. On the other hand, mutations that impair editing out of polymerase α incorporation errors result in cancer displaying a strong mutator phenotype.Įnzymes and reactions in the DNA replication fork. These DNA strand breaks can cause varying forms of chromosome aberrations, contributing to development of cancer that associates with aneuploidy and gross chromosomal rearrangement. Mutations that affect the efficiency of RNA primer removal may result in accumulation of unligated nicks and DNA double-strand breaks. We also discuss studies using mutant mouse models that suggest two distinct cancer etiological mechanisms arising from defects in different steps of Okazaki fragment maturation.  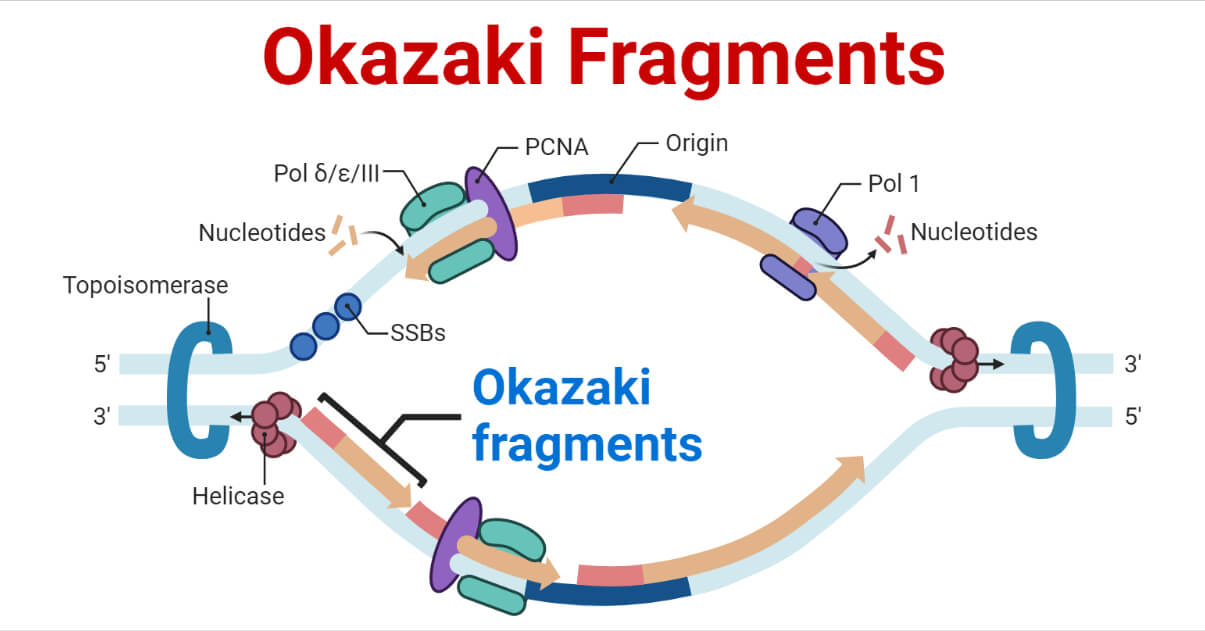
Such protein-protein interactions may be regulated by post-translational modifications. The dynamic interactions of polymerase δ, FEN1 and DNA ligase I with proliferating cell nuclear antigen allow these enzymes to act sequentially during Okazaki fragment maturation. Recent findings reveal that Okazaki fragment maturation is highly coordinated. Here, we summarize the distinct roles of these nucleases in different pathways for removal of RNA/DNA primers. The processing of RNA/DNA primers requires a group of structure-specific nucleases typified by flap endonuclease 1 (FEN1). During this process, primase-synthesized RNA/DNA primers are removed, and Okazaki fragments are joined into an intact lagging strand DNA. Therefore, efficient processing of Okazaki fragments is vital for DNA replication and cell proliferation. Even in yeast, the Okazaki fragment maturation happens approximately a million times during a single round of DNA replication. Completion of lagging strand DNA synthesis requires processing of up to 50 million Okazaki fragments per cell cycle in mammalian cells.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
